Semantic homonym resolution – key to reduce the number of false positive search hits
Many words can have different meanings – also known as “homonyms”. Homonymic terms are often the cause for false positive search hits. How do we use semantic indexing to find what you intended to?
Homonyms in different knowledge areas:
Just take the term “sting” – it could mean a protein named Sting (stimulator of interferon genes), or a known musician, a fictional sword in J. R. R. Tolkien’s The Hobbit and The Lord of the Rings, a 1973 film starring Paul Newman and Robert Redford, a plant disease caused by Belonolaimus longicaudatus, or a structure of an animal to inject venom. In our indexing process we are using the context of each term to establish what is meant.
For example, if other protein terms of the STING protein family are mentioned nearby, if parents or grandparent concepts such as “cyclic-di-GMP binding activity proteins” are also found we are most likely dealing with the protein Sting.
This disambiguation leads, when compared to pure text based indexing, to a reduced but much more precise answer set:
Thus, while PubMed finds 24090 hits and many different meanings when searching for ‘sting’ …
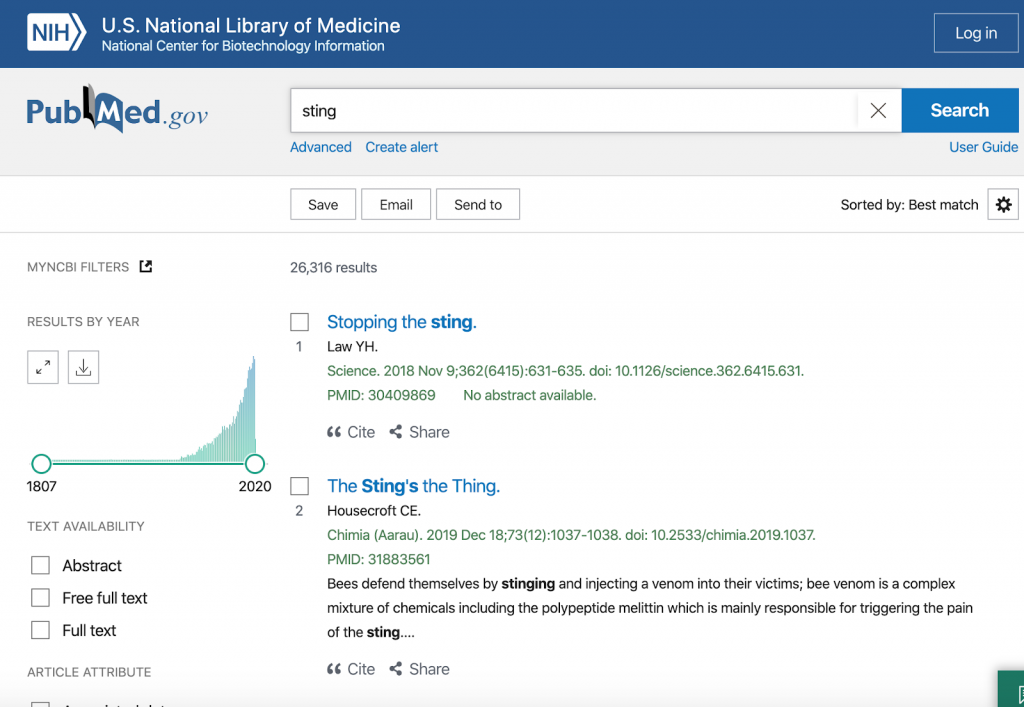
…, searching for ‘sting’ as a protein in our SciWalker will result in only 924 hits in the same document collection:
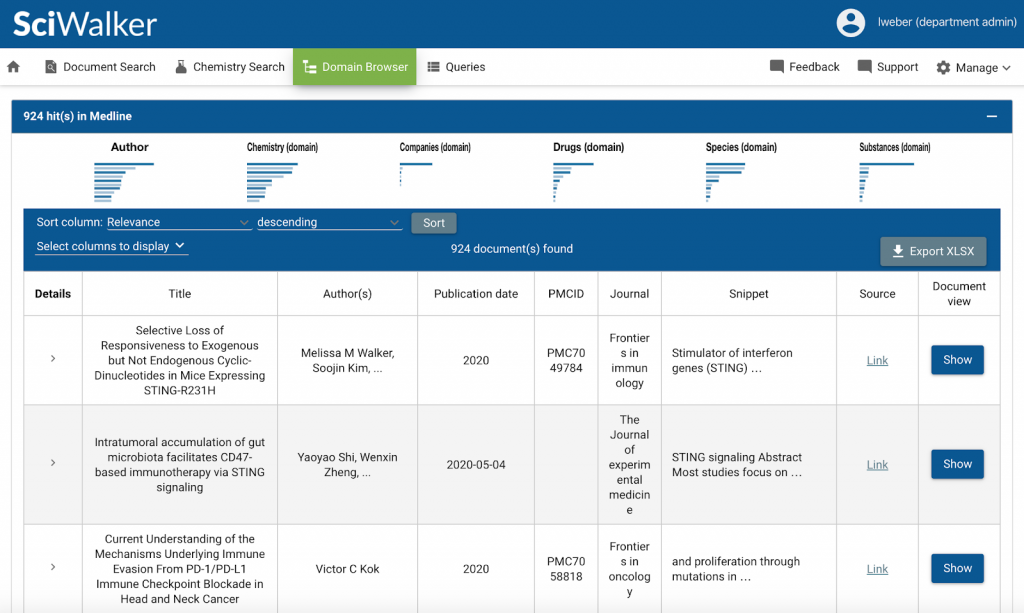
More precise results mean less time is spent browsing through non-relevant documents. It also means that knowledge extraction is so much more reliable and helpful.
Homonyms in the same knowledge area:
Especially the life sciences are full of homonyms, just consider the term “D. melanogaster” – this could mean Drosophila melanogaster (a fly) or Dipoena melanogaster, a spider that actually eats Drosophila melanogaster. To solve this riddle, we are not only looking for terms that are either sister terms or children or parent concepts, but also for the long binominal form somewhere else in the entire document to identify which D. melanogaster is meant.
